Figure 1 from Dominant Mutations in S. cerevisiae PMS1 Identify the Mlh1- Pms1 Endonuclease Active Site and an Exonuclease 1-Independent Mismatch Repair Pathway | Semantic Scholar

PCNA and Msh2-Msh6 activate an Mlh1-Pms1 endonuclease pathway required for Exo1-independent mismatch repair. | Semantic Scholar

Activation of Saccharomyces cerevisiae Mlh1-Pms1 Endonuclease in a Reconstituted Mismatch Repair System* - Journal of Biological Chemistry

Visualization of Eukaryotic DNA Mismatch Repair Reveals Distinct Recognition and Repair Intermediates: Cell

Structure of the MutLα C-terminal domain reveals how Mlh1 contributes to Pms1 endonuclease site | Nature Structural & Molecular Biology
Dominant Mutations in S. cerevisiae PMS1 Identify the Mlh1-Pms1 Endonuclease Active Site and an Exonuclease 1-Independent Mismatch Repair Pathway | PLOS Genetics

The unstructured linker of Mlh1 contains a motif required for endonuclease function which is mutated in cancers | PNAS



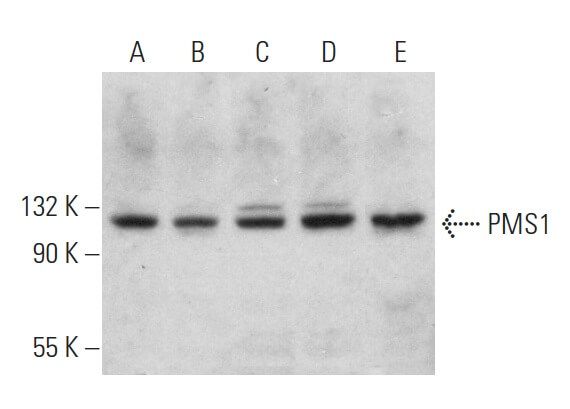

-Western-Blot-NBP2-09011-img0001.jpg)