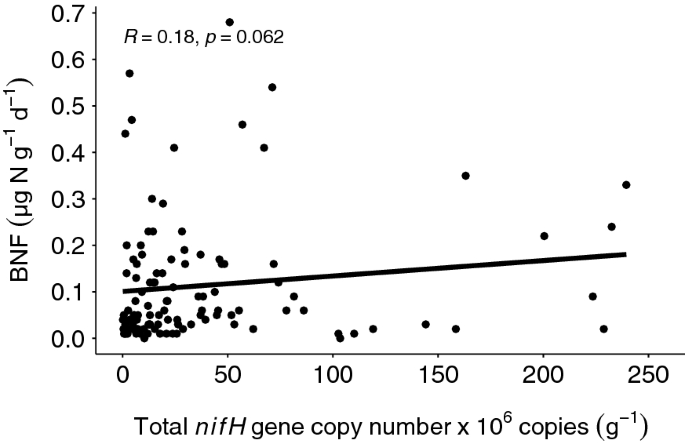
Biological nitrogen fixation and nifH gene abundance in deadwood of 13 different tree species | Biogeochemistry

Nitrogen fixation and nitrogenase (nifH) expression in tropical waters of the eastern North Atlantic | The ISME Journal
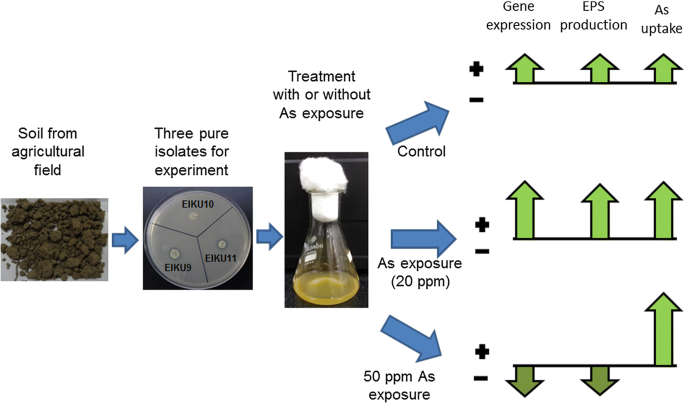
Elevated level of arsenic negatively influences nifH gene expression of isolated soil bacteria in culture condition as well as soil system | SpringerLink

Engineering Nitrogenases for Synthetic Nitrogen Fixation: From Pathway Engineering to Directed Evolution | BioDesign Research

Frontiers | nifH Gene Sequencing Reveals the Effects of Successive Monoculture on the Soil Diazotrophic Microbial Community in Casuarina equisetifolia Plantations
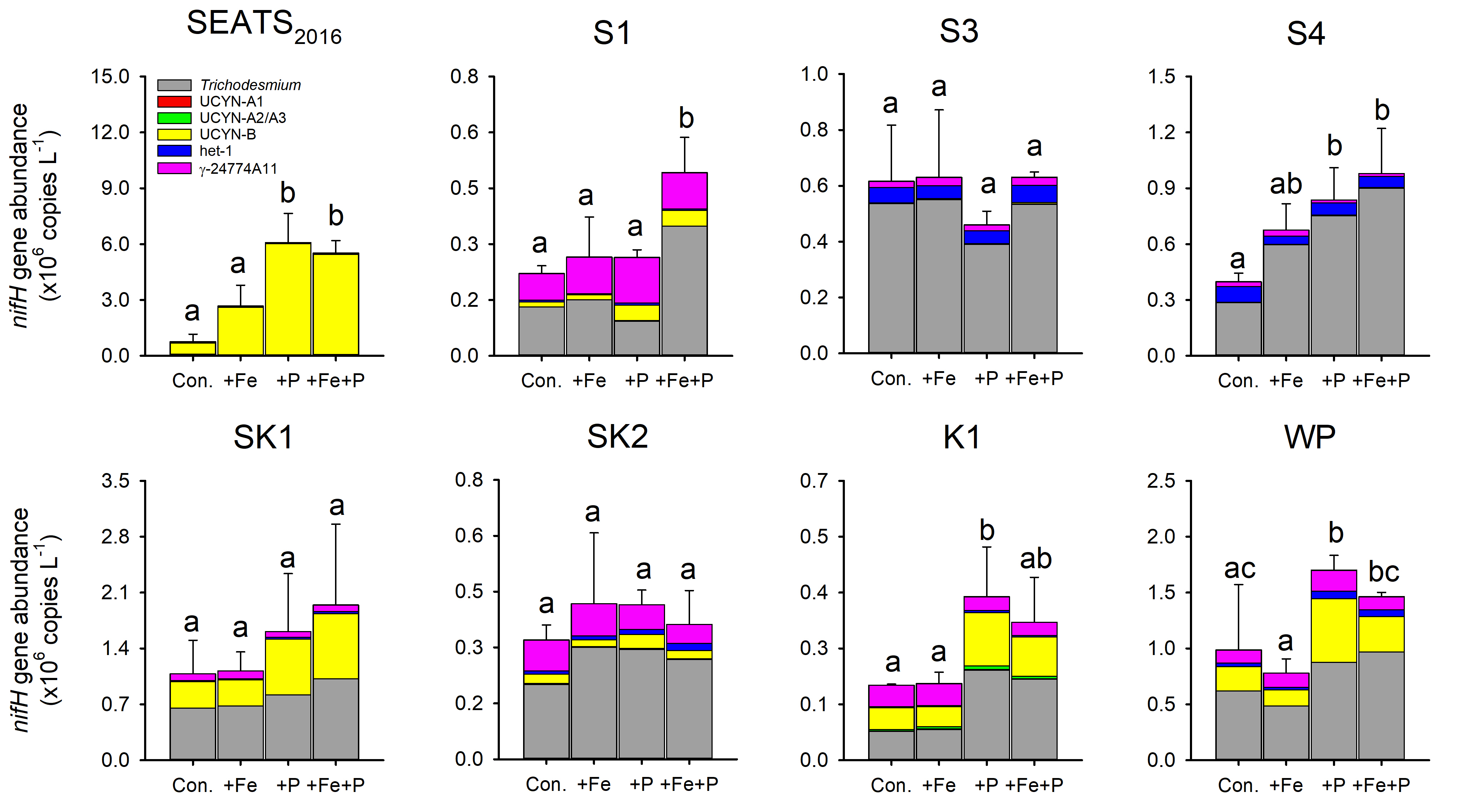
BG - The response of diazotrophs to nutrient amendment in the South China Sea and western North Pacific

Formation of Nitrogenase NifDK Tetramers in the Mitochondria of Saccharomyces cerevisiae | ACS Synthetic Biology
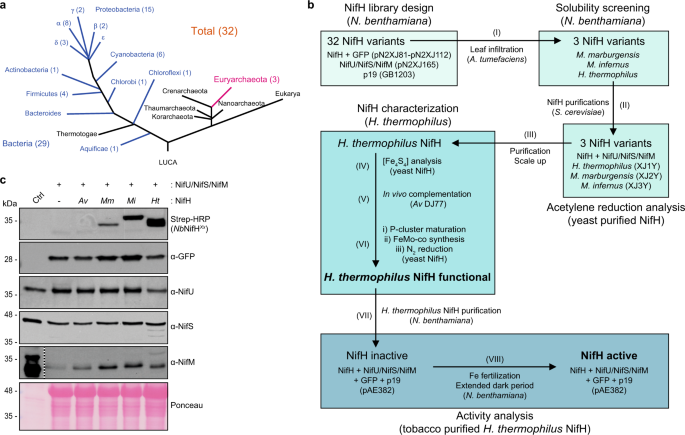
Exploiting genetic diversity and gene synthesis to identify superior nitrogenase NifH protein variants to engineer N2-fixation in plants | Communications Biology
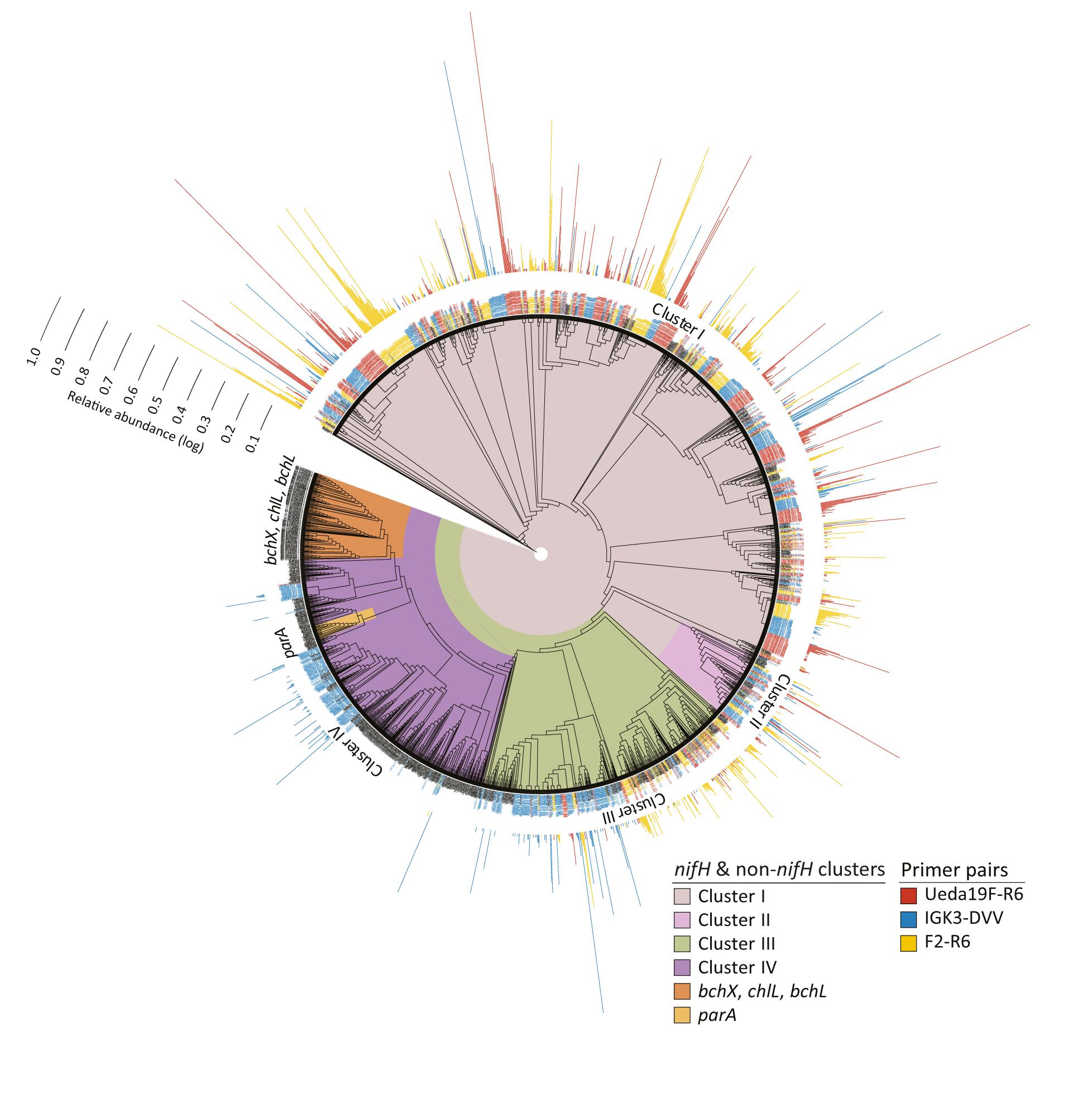
Frontiers | Evaluation of Primers Targeting the Diazotroph Functional Gene and Development of NifMAP – A Bioinformatics Pipeline for Analyzing nifH Amplicon Data

Undervalued Pseudo-nifH Sequences in Public Databases Distort Metagenomic Insights into Biological Nitrogen Fixers | mSphere
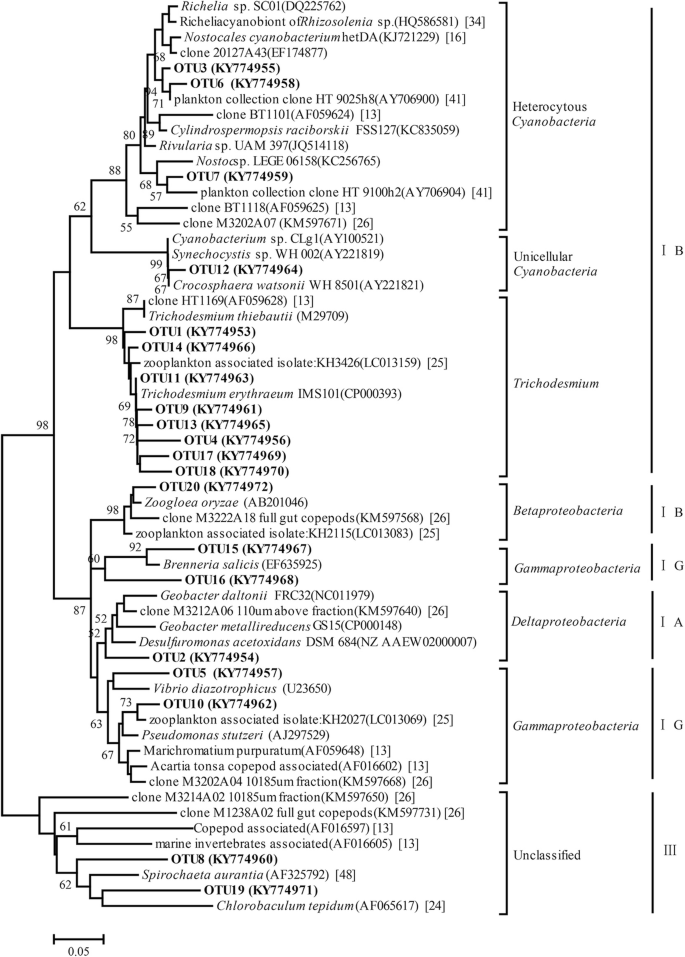
Analysis of nifH DNA and RNA reveals a disproportionate contribution to nitrogenase activities by rare plankton-associated diazotrophs | BMC Microbiology | Full Text

Undervalued Pseudo-nifH Sequences in Public Databases Distort Metagenomic Insights into Biological Nitrogen Fixers | mSphere
Cloning, Prokaryotic Expression and Functional Characterization of NifH Gene from the Associative Nitrogen-Fixing Bacteria Klebsiella Variicola DX120E

The Flora Compositions of Nitrogen-Fixing Bacteria and the Differential Expression of nifH Gene in Pennisetum giganteum z.x.lin Roots
Molecular detection and quantification of nifH gene sequences in the rhizosphere of sorghum (Sorghum bicolor) sown with two leve

Characterization of a nitrogen-fixation (nif) gene cluster from Anabaena azollae 1a shows that closely related cyanobacteria have highly variable but structured intergenic regions | Microbiology Society

Seasonal and long-term effects of nutrient additions and liming on the nifH gene in cerrado soils under native vegetation - ScienceDirect
Temporal Dynamics of Abundance and Composition of Nitrogen-Fixing Communities across Agricultural Soils | PLOS ONE
![Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors](https://pub.mdpi-res.com/genes/genes-13-01428/article_deploy/html/images/genes-13-01428-g001.png?1660208416)
Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors